RNA Splicing and MDS
The most common mutations found in MDS occur in genes involved in RNA splicing (including SF3B1, SRSF2, U2AF1, and ZRSR2) and epigenetic modification (including TET2, ASXL1, and DNMT3A). The identification of spliceosome mutations in approximately half of all patients with MDS implicates abnormalities of RNA splicing.
The most common mutations found in MDS occur in genes involved in RNA splicing (including SF3B1, SRSF2, U2AF1, and ZRSR2) and epigenetic modification (including TET2, ASXL1, and DNMT3A). The identification of spliceosome mutations in approximately half of all patients with MDS implicates abnormalities of RNA splicing.
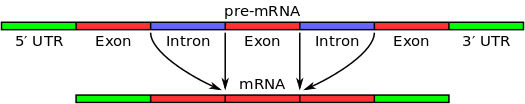
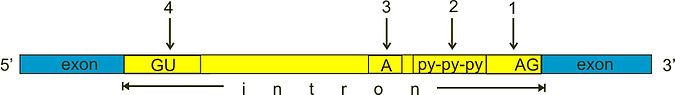
The first stage in RNA splicing is recognition by the spliceosome of splice sites between introns and exons. Key to this process are short sequence motifs. These include the 5' and 3' splice sites (typically a GU and AG sequence respectively); the branch point sequence (which contains a conserved adenosine important to intron removal); and the polypyrimidine tract (which is thought to recruit factors to the branch point sequence and 3' splice site). These sequence motifs are represented in the illustration below:
The U1 snRNP recognizes and binds to the 5' splice site. The branch point sequence is identified and bound by the branch-point-binding protein (BBP). The 3' splice site and polypyrimidine tract are recognized and bound by two specific components of a protein complex called U2 auxiliary factor (U2AF): U2AF35 and U2AF65 respectively.
The figure above shows the binding of SR proteins to splicing enhancer sites, which promotes the binding of U1 snRNP to the 5' splice site, and U2AF protein to the polypyrimidine tract and 3' splice site. The U2AF1 heterodimer is generally accepted to play a vital role in defining functional 3' splice sites in pre-mRNA splicing.